Bioinformatics and Unsupervised Machine Learning Tutorial
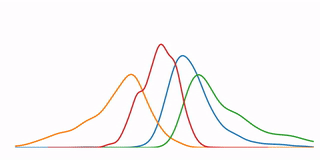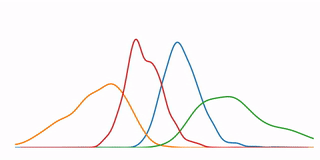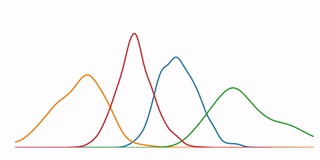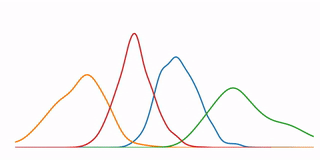
A tutorial that demonstrates some high-level concepts in bioinformatics, undirected machine learning, and data visualization. In particular, it performs a K-means clustering analysis on microarray data from 6153 genes at 7 time points, and demonstrates several methods for visualizing the resulting data.
The code of this notebook is all written in Python 3, and a copy of this notebook is available at My GitHub.
Brownian Crystal Generator

A diffusion limited aggregation program written in python as a personal project in complexity science. It's a cellular automata where particles take random walks across a canvas, adding themselves to a seed crystal when they collide with it. Data about the particles is logged with each new addition, allowing a statistical analysis of the simulation. It's also highly customizable, from colors, to the movement of the particles, and even allows for real-time rendering.
A Simple Supervised Neural Network
A small and light (core code less than 20 lines of python) neural net constructed as a teaching example in Machine Learning. It loads an input and output dataset, then loops 100000 times, backprogpagating error correction each loop to arrive at a trained net to which you can present new situations.
This Website

This website was written by me as well. Though I don't pretend to have a mastery of HTML or CSS, I think it's pretty okay.
It's under an MIT licence, so if you want to borrow some of my code that's cool, just give attributation.